Medical research into the complex and detailed field of cell function, genetics, and related diseases use antibodies to detect where proteins are in the cell and their number. Proteins are biomolecules that translate information from an organism’s genes into the structure, function, and regulation of its tissues and organs.
Researchers often purchase commercially available antibodies for their research, antibodies specifically designed to detect specific proteins. It turns out however that it’s not entirely unknown in the science community that these antibodies may not always be as accurate as hoped, leading to research results that themselves may not be accurate or that can be validated through subsequent research.
Research at the Montreal Neurological Institute highlighted the problem and devised a procedure other labs can use to validate their research antibodies to ensure accurate research results.
Peter McPherson (PhD., FRSC) professor at McGill University, and researcher at the Montreal Neurological Institute and Hospital (the Neuro)
Genetic mutations can cause protein imbalances or malfunctions, leading to human disease, therefore locating protiens, their type and number is critical in medical research. If the antibodies are not accurate, the whole research result may be called into question.
For some time researchers have suspected that the commercial antibodies may not be entirely accurate, but there are a couple of factors playing into their continued use “as is”. These are the time and cost of the validation process which detracts from the time and funding for the desired research project. Not validating the antibodies beforehand however may compromise the results of the much more time consuming and expensive research and thus be a false economy.
McPherson’s team chose a specific protein product of a gene, C9ORF72. Mutations of this are the major genetic cause of amyotrophic lateral sclerosis (ALS) and frontotemporal dementia. They tested 16 antibodies advertised as specific to this protein, but found only one of the 16 accurately detected C9ORF72 in immunofluorescence, a technique in which antibodies stain proteins so they can be seen under a microscope. The others either didn’t detect the protein or additionally detected other unrelated proteins as well. Using other techniques, two other antibodies detected the protein, but the successful antibodies have not yet been used in published research, whereas the unsuccessful ones have been used in several studies, calling the results into question.
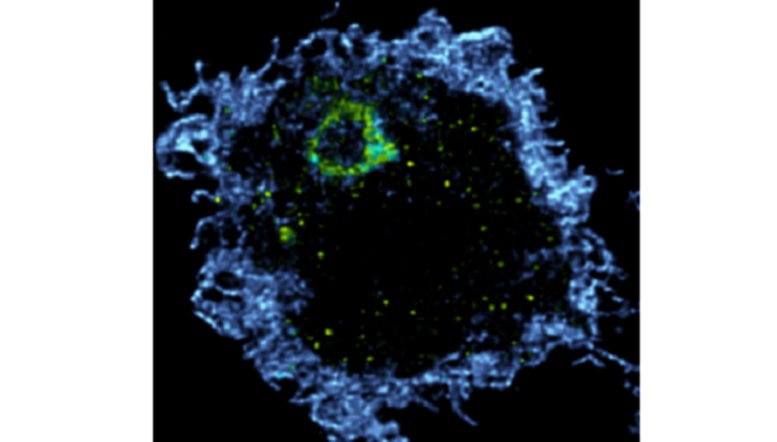
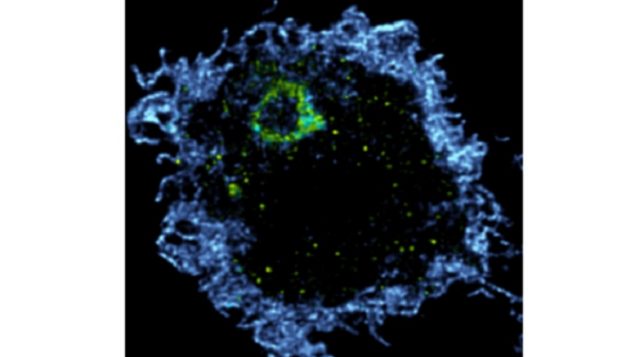
This microscopy image shows a human macrophage stained for C9ORF72.. Mutations of C9ORF72 seem to be directly associated with ALS and the activity of macrophages which are the immune system ‘claeaners” Macrophages ingest pathogens or debris in large phagosomes. As showed here, C9ORF72 localizes on a phagosome (yellow). Blue is the actin cytoskeleton. (LaFlamme et al- McGill- The Nuero)
Professor McPherson says the commercial suppliers are working to improve their products, but for now researchers can use the relatively simple template his team created which other labs can follow to validate their antibodies before proceding with the main thrust of research.
Quoted in a McGill University press release he said, “The procedures we use are not revolutionary, and in fact this makes our approach widely applicable to any laboratory skilled in the art, yet to my knowledge this is one of the first papers to describe a streamlined process for antibody validation. A large part of the reproducibility crisis is because of poor antibody validation. We owe it to funders and patients to do better”.
The research was published in the open access journal eLife on Oct. 15, 2019, https://elifesciences.org/articles/48363
This work was supported by a grant from the Motor Neurone Disease Association (UK), The ALS Association (USA), The ALS Society of Canada and by an ALS Canada-Brain Canada Arthur J. Hudson Translational Team Grant. It was completed as part of the ALS-Reproducible Antibody Platform, a project designed to test antibodies against the protein products of ALS disease gene






For reasons beyond our control, and for an undetermined period of time, our comment section is now closed. However, our social networks remain open to your contributions.